Poly(A) Polymerase, Yeast
$0.00 – $453.20
Poly(A) Polymerase from Yeast works more efficiently than E. coli Poly(A) Polymerase for RNA oligonucleotide-labeling and poly(A) tailing.
| SKU | OPTIONS | Price | Quantity | ||
|---|---|---|---|---|---|
| PAPY-30 | 120,000 units (600 U/ul) | $141.63 | |||
| PAPY-40 | 240,000 units (600 U/ul) | $226.60 | |||
| PAPY-50 | 600,000 units (600 U/ul) | $453.20 | |||
| PAPY-OEM | Any Size | Please inquire |
- Description
- Additional information
- Documents
- Q&A
Description
Description:
Poly(A) Polymerase catalyzes the template independent of the addition of AMP from ATP to the 3′-end of RNA. Poly(A) works more competently than E. coli poly(A) polymerase for RNA oligonucleotide-labeling and poly(A) tailing. Less incubation time is required for the yeast enzyme. This enzyme labels both long and short substrates. Poly(A) polymerase preferentially labels longer RNA-molecules whereas short RNA-molecules are labeled more efficiently by T4 RNA ligase. The reaction requires Mn2+ or Mg2+, ATP as substrates, and any RNA containing 3′-hydroxyl termini as primers. Longer RNA molecules are somewhat better primers than short oligomers. Substitution of cordycepin 5′-triphosphate (3′-dATP) for ATP results in the addition of a single 3′-dA residue to the ends of the RNA, a useful technique for labeling RNA at the 3′-end.
Applications:
- Labeling the 3′-ends of RNA with ATP or cordycepin
- Poly(A) tailing of RNA for cloning or affinity purification
- Preparing a priming site for cDNA synthesis using oligo-dT
- Enhancing translation of RNA transferred into eukaryotic cells
Source:
An E. coli strain that carries the cloned Poly(A) Polymerase gene from Saccharomyces cerevisiae
Specific Activity: >20,000 U/mg
Unit Definition:
One unit is the amount of enzyme which incorporates 1 pmol AMP into acid-insoluble material at 37°C in 1 min.
5x Poly(A) Polymerase Reaction Buffer:
100 mM Tris-HCl, pH 7.0, 3.0 mM MnCl2, 0.1 mM EDTA, 1 mM DTT, 500 µg/ml Acetylated BSA, 50% Glycerol.
Storage Buffer:
20 mM Tris-HCl (pH 8.0), 50 mM KCl, 0.5 mM DTT, 50% Glycerol
Assay Conditions:
1x Poly(A) Polymerase Reaction Buffer, 1 mM rATP and 500 ng 5′-FAM labeled poly A 20-mer RNA in a 20 µl reaction. After incubation at 37°C for 10 min, acid insoluble radioactivity is determined either by gel electrophoresis or with an automated capillary DNA sequencer. In this assay 5 units of enzyme add approximatley 60 to 80 adenosines to the RNA primer. In these conditons 20 units of enzyme will deplete the rATP.
Heat Inactivation: 65°C for 20 minutes
Recommended Storage Condition: -20ºC
References:
1. Sippel, A. E. (1973) Eur. J. Biochem. 37, 31-40.
2. Edmonds, M. (1982) in The Enzymes, 3rd edition, ed. P. D. Boyer (Academic Press, New York) 15, 217-244.
3. Gething, M. J., Bye, J., Skehel, J. and Waterfield, M. (1980) Nature 287, 301-306.
4. Sano, H. and Feix, G. (1976) Eur. J. Biochem. 71, 577-583.
Additional information
| OPTIONS | 120,000 units (600 U/ul), 240,000 units (600 U/ul), 600,000 units (600 U/ul), Any Size |
|---|
Manual & Protocols
MSDS & Certificates
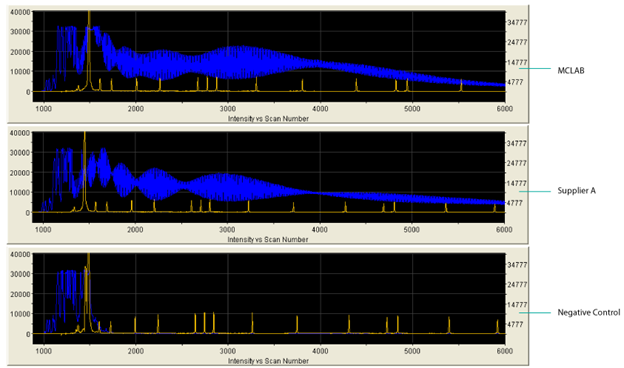
Figure Poly (A) Tailing FAM-Labeled RNA
Performance of Poly (A) Polymerase, Yeast either from MCLAB or another supplier was compared with standard protocol. Paralleled negative control was run without enzyme. The results are analyzed by ABI 3730xl Data Analyzer.